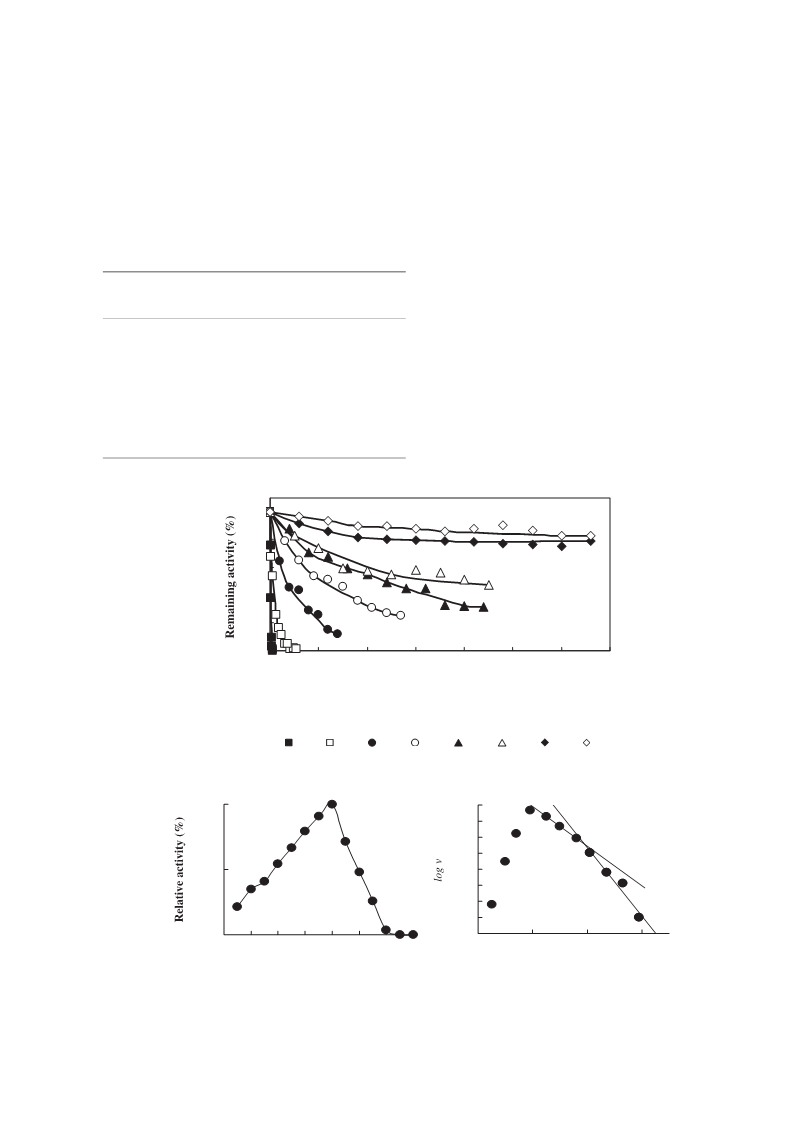
Malate Dehydrogenase from Flavobacterium frigidimaris KUC-1
2153
Biochem. Z., 336, 371–379 (1962).
1: enzymological characteristics and functional proper-
ties. Biochem. Biophys. Res. Commun., 298, 632–637
(2002).
2) Murphey, W. H., Barnaby, C., Lin, F. J., and Kaplan,
N. O., Malate dehydrogenases. II. Purification and
properties of Bacillus subtilis, Bacillus stearothermo-
philus, and Escherichia coli malate dehydrogenases.
J. Biol. Chem., 242, 1548–1559 (1967).
3) Phizackerley, P. J., and Francis, M. J., Cofactor require-
ments of the L-malate dehydrogenase of Pseudomonas
ovalis Chester. Biochem. J., 101, 524–535 (1966).
4) Shrago, E., and Falcone, A. B., Purification and proper-
ties of human-erythrocyte malic dehydrogenase. Bio-
chim. Biophys. Acta, 73, 7–16 (1963).
5) Weimberg, R., Effect of sodium chloride on the activity
of a soluble malate dehydrogenase from pea seeds.
J. Biol. Chem., 242, 3000–3006 (1967).
6) Yoshida, A., Purification and chemical characterization
of malate dehydrogenase of Bacillus subtilis. J. Biol.
Chem., 240, 1113–1117 (1965).
19) Kazuoka, T., Takigawa, S., Arakawa, N., Hizukuri, Y.,
Muraoka, I., Oikawa, T., and Soda, K., Novel psychro-
philic and thermolabile L-threonine dehydrogenase from
psychrophilic Cytophaga sp. strain KUC-1, J. Bacteriol.,
185, 4483–4489 (2003).
20) Oikawa, T., Yamanaka, K., Kazuoka, T., Kanzawa, N.,
and Soda, K., Psychrophilic valine dehydrogenase of the
antarctic psychrophile, Cytophaga sp. KUC-1: purifica-
tion, molecular characterization and expression. Eur. J.
Biochem., 268, 4375–4383 (2001).
21) Velick, S. F., and Vavra, J., A kinetic and equilibrium
analysis of the glutamate oxalacetate transaminase
mechnism. J. Biol. Chem., 237, 2109–2122 (1962).
22) Esaki, N., Shimoi, H., Nakajima, N., Ohshima, T.,
Tanaka, H., and Soda, K., Enzymatic in situ determi-
nation of stereospecificity of NAD-dependent dehydro-
genases. J. Biol. Chem., 264, 9750–9752 (1989).
23) Bradford, M. M., A rapid and sensitive method for the
quantitation of microgram quantities of protein utilizing
the principle of protein-dye binding. Anal. Biochem., 72,
248–254 (1976).
7) Tamegai, H., Li, L., Masui, N., and Kato, C., A
denitrifying bacterium from the deep sea at 11,000-m
depth. Extremophiles, 1, 207–211 (1997).
8) Kraegeloh, A., and Kunte, H. J., Novel insights into the
role of potassium for osmoregulation in Halomonas
elongata. Extremophiles, 6, 453–462 (2002).
9) Grant, W. D., Gemmell, R. T., and McGenity, T. J.,
Halobacteria: the evidence for longevity. Extremophiles,
2, 279–287 (1998).
24) Tulchin, N., Ornstein, L., and Davis, B. J., A microgel
system for disc electrophoresis. Anal. Biochem., 72,
485–490 (1976).
10) Breuil, C., and Kushner, D. J., Lipase and esterase
formation by psychrophilic and mesophilic Acineto-
bacter species. Can. J. Microbiol., 21, 423–433 (1975).
11) Zakaria, M. M., Ashiuchi, M., Yamamoto, S., and Yagi,
T., Optimization for beta-mannanase production of a
psychrophilic bacterium, Flavobacterium sp. Biosci.
Biotechnol. Biochem., 62, 655–660 (1998).
25) Laemmli, U. K., Cleavage of structural proteins during
the assembly of the head of bacteriophage T4. Nature,
227, 680–685 (1970).
26) Oikawa, T., Tsukagawa, Y., and Soda, K., Endo-ꢁ-
glucanase secreted by a psychrotrophic yeast: purifica-
tion and characterization. Biosci. Biotechnol. Biochem.,
62, 1751–1756 (1998).
12) Feller, G., Zekhnini, Z., Lamotte-Brasseur, J., and
Gerday, C., Enzymes from cold-adapted microorgan-
isms: the class C beta-lactamase from the antarctic
psychrophile Psychrobacter immobilis A5. Eur. J. Bio-
chem., 244, 186–191 (1997).
13) Bruni, V., Gugliandolo, C., Maugeri, T., and Allegra, A.,
Psychrotrophic bacteria from a coastal station in the
Ross Sea (Terra Nova Bay, Frigidimaris). New Micro-
biol., 22, 357–363 (1999).
14) Kobori, H., Sullivan, C. W., and Shizuya, H., Heat-labile
alkaline phosphatase from Antarctic bacteria: rapid 50
end-labeling of nucleic acids. Proc. Natl. Acad. Sci.
U.S.A., 81, 6691–6695 (1984).
15) Vckovski, V., Schlatter, D., and Zuber, H., Structure and
function of L-lactate dehydrogenase from thermophilic,
mesophilic and psychrophilic bacteria. IX. Identification,
isolation, and nucleotide sequence of two L-lactate
dehydrogenase genes of the psychrophilic bacterium
Bacillus psychrosaccharolyticus. Biol. Chem. Hoppe-
Seyler, 371, 103–110 (1990).
27) Tyagi, A. K., Siddiqui, F. A., and Venkitasubramanian,
T. A., Studies on the purification and characterization of
malate dehydrogenase from Mycobacterium phlei. Bio-
chim. Biophys. Acta, 485, 255–267 (1977).
28) Kim, S. Y., Hwang, K. Y., Kim, S. H., Sung, H. C., Han,
Y. S., and Cho, Y., Structural basis for cold adaptation.
Sequence, biochemical properties, and crystal structure
of malate dehydrogenase from a psychrophile Aquaspir-
illium arcticum. J. Biol. Chem., 274, 11761–11767
(1999).
29) Birktoft, J. J., Fernley, R. T., Bradshaw, R. A., and
Banaszak, L. J., Amino acid sequence homology among
the 2-hydroxy acid dehydrogenases: mitochondrial and
cytoplasmic malate dehydrogenases form a homologous
system with lactate dehydrogenase. Proc. Natl. Acad.
Sci. U.S.A., 79, 6166–6170 (1997).
30) Langelandsvik, A. S., Steen, I. H., Birkeland, N. K., and
Lien, T., Properties and primary structure of a thermo-
stable L-malate dehydrogenase from Archaeoglobus
fulgidus. Arch. Microbiol., 168, 59–67 (1997).
16) Gianese, G., Bossa, F., and Pascarella, S., Comparative
structural analysis of psychrophilic and meso- and
thermophilic enzymes. Proteins, 47, 236–249 (2002).
17) Nogi, Y., Soda, K., and Oikawa, T., Flavobacterium
frigidimaris sp. nov., isolated from Antarctic seawater.
Sys. Appl. Microbiol., 28, 310–315 (2005).
18) Yamanaka, Y., Kazuoka, T., Yoshida, M., Yamanaka,
K., Oikawa, T., and Soda, K., Thermostable aldehyde
dehydrogenase from psychrophile, Cytophaga sp. KUC-
31) Nicholls, D. J., Miller, J., Scawen, M. D., Clarke, A. R.,
Holbrook, J. J., Atkinson, T., and Goward, C. R., The
importance of arginine 102 for the substrate specificity
of Escherichia coli malate dehydrogenase. Biochem.
Biophys. Res. Commun., 189, 1057–1062 (1992).
32) Charnock, C., Refseth, U. H., and Sirevag, R., Malate
dehydrogenase from Chlorobium vibrioforme, Chloro-
bium tepidum, and Heliobacterium gestii: purification,
characterization, and investigation of dinucleotide bind-

 Oikawa, Tadao
Oikawa, Tadao
 Yamamoto, Noriko
Yamamoto, Noriko
 Shimoke, Koji
Shimoke, Koji
 Uesato, Shinichi
Uesato, Shinichi
 Ikeuchi, Toshihiko
Ikeuchi, Toshihiko
 Fujioka, Toru
Fujioka, Toru