4
2
M. Yamada et al. / Journal of Molecular Catalysis B: Enzymatic 105 (2014) 41–48
O
development solution described below in Section 2.4. The iso-
OH
lated strains were incubated in a test tube (16 × 1.5 cm diameter)
HO
◦
containing 5 ml of the screening medium at 30 C for 3 days. Sub-
Glycolate
O
sequently, cell-free extract was prepared by disrupting the cells at
O
◦
HO
Ethyleneglycol
O
below 5 C for 8 min by a Multi-beads shocker (Yasui Kikai, Osaka,
OH
Glycolaldehyde
OH
HO
Glyoxylate
Japan). The strain exhibiting high activity toward glycolate but not
toward glyoxylate was selected and used in this study.
O
O
Glyoxal
2.3. Identification of the isolated strain
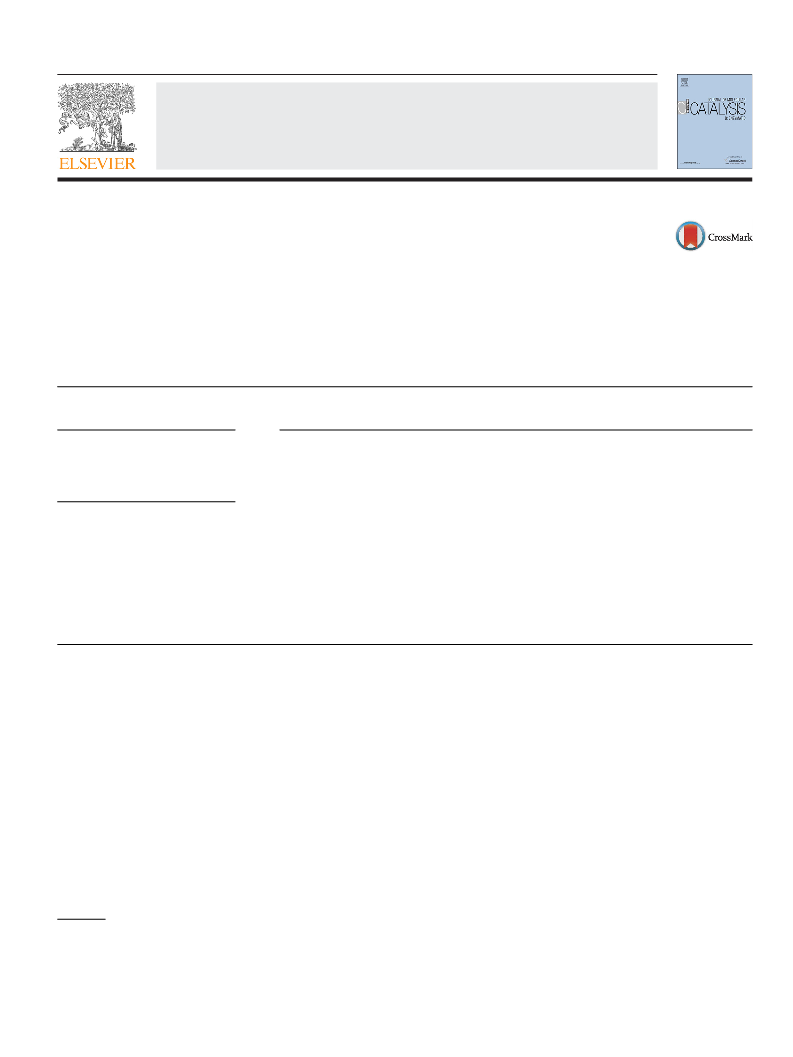
Fig. 1. Pathways of enzymatic synthesis of glyoxylate from ethylene glycol.
Identification of the newly isolated strain was performed at
NCIMB Japan Co., Ltd. (Shizuoka, Japan).
pathway, the conversion of ethylene glycol to glycolate, it was
reported that ethylene glycol-oxidizing microorganisms such as
Acetobactor [10], Gluconobacter [11], and Hansenula [12] accumu-
lated glycolate in media with ethylene glycol. Furthermore, Kataoka
et al. screened and isolated microorganisms, Pichia naganishii AKU
2.4. Cultivation of the isolated strain
The isolated strain was incubated in 5 ml of the 1,2-propanediol
medium containing 1% 1,2-propanediol, 0.2% (NH4)2SO4, 0.1%
K2HPO4, 0.1% NaH2PO4·2H2O, 0.05% MgSO4·7H2O, 0.02%
CaCl2·2H2O, 0.5% corn steep liquor, and 0.05% yeast extract
4
267 and Rhodotorula sp. 3Pr-126, which produced high concen-
trations of glycolate in media containing high concentrations of
ethylene glycol [13]. It would be assumed that these microor-
ganisms have enzymes which catalyze the conversion of ethylene
glycol into glycolate via glycolaldehyde, although those enzymes
have not been purified and characterized. In the conversion of gly-
colate into glyoxylate, glycolate dehydrogenase from Trichoderma
harzianum AIU 353 [14] and glycerol oxidase from A. japonicus [5] to
catalyze the conversion reaction. In addition, it is well known that
various glycolate oxidases (EC 1.1.3.15) from many green plants
and mammalian livers catalyze this conversion reaction. The gly-
colate oxidases have been isolated from a variety of sources, such
as spinach [15], sugar beet [16], pea [17], pumpkin [18], cucumber
cotyledons [19], lettuce [20], tobacco [21], rat liver [22], chicken
liver [23], and human liver [24]. The X-ray crystal structures of
the glycolate oxidases from spinach [25] and human liver [26]
have been determined. Regarding microorganisms, genes encod-
ing the subunits of glycolate oxidase (glc DEF) from Escherichia coli
have been identified, but their enzymatic properties have not been
characterized in detail [27]. Among these glycolate oxidases, the
glycolate oxidase from spinach has been used for the enzymatic
production of glyoxylate [1–3]. However, the microbial glycolate
oxidase was not reported. We therefore screened for an oxidase
that would catalyze the conversion of glycolate into glyoxylate but
not the conversion of glyoxylate into oxalate.
◦
(pH 6.0) at 30 C for 2 days with shaking (120 strokes/min).
The culture (1 ml) was inoculated into a 500-ml culture flask
containing 150 ml of the 1,2-propanediol medium, then incubated
◦
at 30 C for 3 days with shaking. A volume of 40 ml of the second
culture was transferred into a 3 l culture flask containing 2 l of the
1,2-propanediol medium, then cultivated at 30 C for 3 days with
shaking.
◦
2.5. Glyoxylate production from glycolate by resting-cell reaction
Approximately 13 g (wet cell weight) of resting cells from
the isolated strain harvested after 3 days of cultivation in 1,2-
propanediol medium were incubated with 1.0 M glycolate in 15 ml
◦
of 0.1 M potassium phosphate buffer (pH 6.0) at 20 C for 14 days.
Glyoxylate in the reaction mixture was analyzed by HPLC with
an ULTRON PS-80H column (Shinwa Chemical Industries, Tokyo,
Japan). The elution was carried out at a flow rate of 1.0 ml per min at
◦
60 C with perchloric acid solution (pH 2.1) for 20 min. The elution
peaks of glycolate, glyoxylate, and oxalate were eluted at 8.8 min,
7.2 min, and 5.3 min, respectively, under the same experimental
conditions.
2.6. Enzyme activity assay
The present paper describes the identification of the isolated
strain and certain properties of oxidase using the purified enzyme.
Oxidase activity was assayed by measuring the forma-
◦
tion rate of hydrogen peroxide at 30 C. The reaction mixture
was composed of 1 ml of a solution containing 50 mol of
glycolate, 0.6 mol of 4-aminoantipyrine, 1.94 mol of N-ethyl-N-
(2-hydroxy-3-sulfopropyl)-3-methylaniline sodium salt dihydrate,
2
. Materials and methods
2.1. Chemicals
6
.7 units of peroxidase, 0.1 mmol of potassium phosphate (pH 5.5),
Sodium glycolate, glyoxylate monohydrate, and 3-methyl-2-
and an appropriate amount of enzyme solution. The formation
of hydrogen peroxidase was spectrophotometrically measured by
following the increase in the absorbance at 555 nm. One unit of
enzyme activity was determined as the amount of enzyme catalyz-
ing the formation of one micromole of hydrogen peroxidase per
minute under the conditions above.
benzothiazolinone hydrazine hydrochloride (MBTH) were pur-
chased from Wako Pure Chemical Industries (Osaka, Japan).
Horseradish peroxidase (EC 1.11.1.7) was obtained from Amano
Enzyme (Nagoya, Japan). All other chemicals used were the highest
grade that is commercially available.
2.2. Isolation of microorganisms
2.7. Purification of the enzyme
◦
The enrichment culture was carried out three times using a
All the procedures were performed at 5–10 C using potassium
screening medium containing 5% 1,2-propanediol, which has a
similar structure to glycolate, 0.2% (NH ) SO , 0.1% K HPO , 0.1%
phosphate buffer (pH 6.0). The cells from 20 l of culture broth
(128 g of wet weight) were disrupted with glass beads in 10 mM
buffer solution by a Multi-beads shocker, and the supernatant (1.5 l)
collected by centrifugation at 10,000 × g for 30 min was used as
a crude enzyme solution. Then, solid ammonium sulfate (363 g)
was added to the crude enzyme solution to reach 40% saturation
4
2
4
2
4
NaH PO ·2H O, 0.02% MgSO ·7H O, 0.01% CaCl ·2H O, and 0.05%
2
4
2
4
2
2
2
yeast extract (pH 6.0). The microorganisms were cultivated on an
◦
agar plate of the screening medium at 30 C for 2–3 days. The
formation of hydrogen peroxide was detected by using the color

 Yamada, Miwa
Yamada, Miwa
 Higashiyama, Takanori
Higashiyama, Takanori
 Kishino, Shigenobu
Kishino, Shigenobu
 Kataoka, Michihiko
Kataoka, Michihiko
 Ogawa, Jun
Ogawa, Jun
 Shimizu, Sakayu
Shimizu, Sakayu
 Isobe, Kimiyasu
Isobe, Kimiyasu